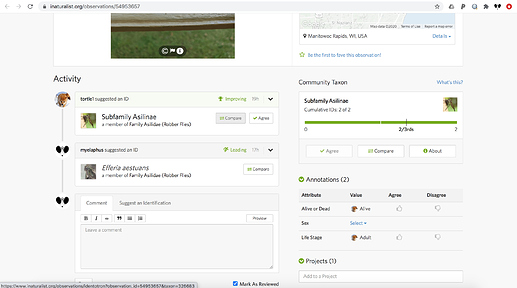
i looked at the above issue related to num_identification_disagreements with fresh eyes this morning, and here’s what i see when i click on “About” in the community taxon summary for the observation mentioned above (https://www.inaturalist.org/observations/54972306):
note that at the subfamily level there are no disagreements because all the IDs are for Asilinae. in other words, even though the IDs disagree with each other, they don’t disagree with the community taxon, and so num_identification_disagreements probably represents disagreements with the community taxon. (that’s why it’s zero.)
i think the most_agree, some_agree, etc. also operate the same way. so none of these seem very useful in the context of what you’re asking for.
it still would be possible to loop through the active identifications, counting (distinct) taxa that, compared to the observation taxon, are:
- (the same as) the observation taxon
- ancestor taxa
- descendant taxa
- other taxa
… and some sort of metric based on that that might be the most informative in this context. i’m still trying to figure out a way to represent that that ideally does not take up 4 columns though. i also don’t want to get too far away from the basic information that the API response provides, since https://jumear.github.io/stirfry/iNatAPIv1_observations.html is supposed to be a simple page that reformats the endpoint response to be more human-readable, not a page that does a lot of extra stuff on top of the response.
so if anyone has any suggestions for best approach, feel free to share.
…
UPDATE:
no one offered any suggestions, and i couldn’t think of a better way. so I went ahead and updated the page noted above to add some extra columns for ID count and (count of) ID taxa in various categories. so https://jumear.github.io/stirfry/iNatAPIv1_observations.html?exact_taxon_id=326683&place_id=1&per_page=200 should pull back all US Asilinae, and then to exclude observations whose IDs don’t agree, you can skip past the ones that have values > 0 in either the “id taxa @ desc”(endant) or “id taxa @ other” columns.
so, for example, here’s what i see in that page for US Asilinae right now:
note that observation 55154044 (https://www.inaturalist.org/observations/55154044) is shown as having 2 identification taxa that are descendants of the observation taxon. since there are also 0 ID taxa at the observation taxon, you can interpret that as meaning there are lower-level IDs that disagree with each other, and that’s in fact what we see in the observation itself:
hope that all makes sense… if else has any additional thoughts or suggestions, let me know.